Overview
If you’re completing a review that looks only at the effects of health interventions through Covidence and want to only consider research papers reporting on Randomised Controlled Trials (RCTs), consider using this feature.
All eligible references imported to the review will be run through the Cochrane RCT classifier and tagged with either “Possible RCT” or “Not RCT”, depending on the prediction made by the classifier:
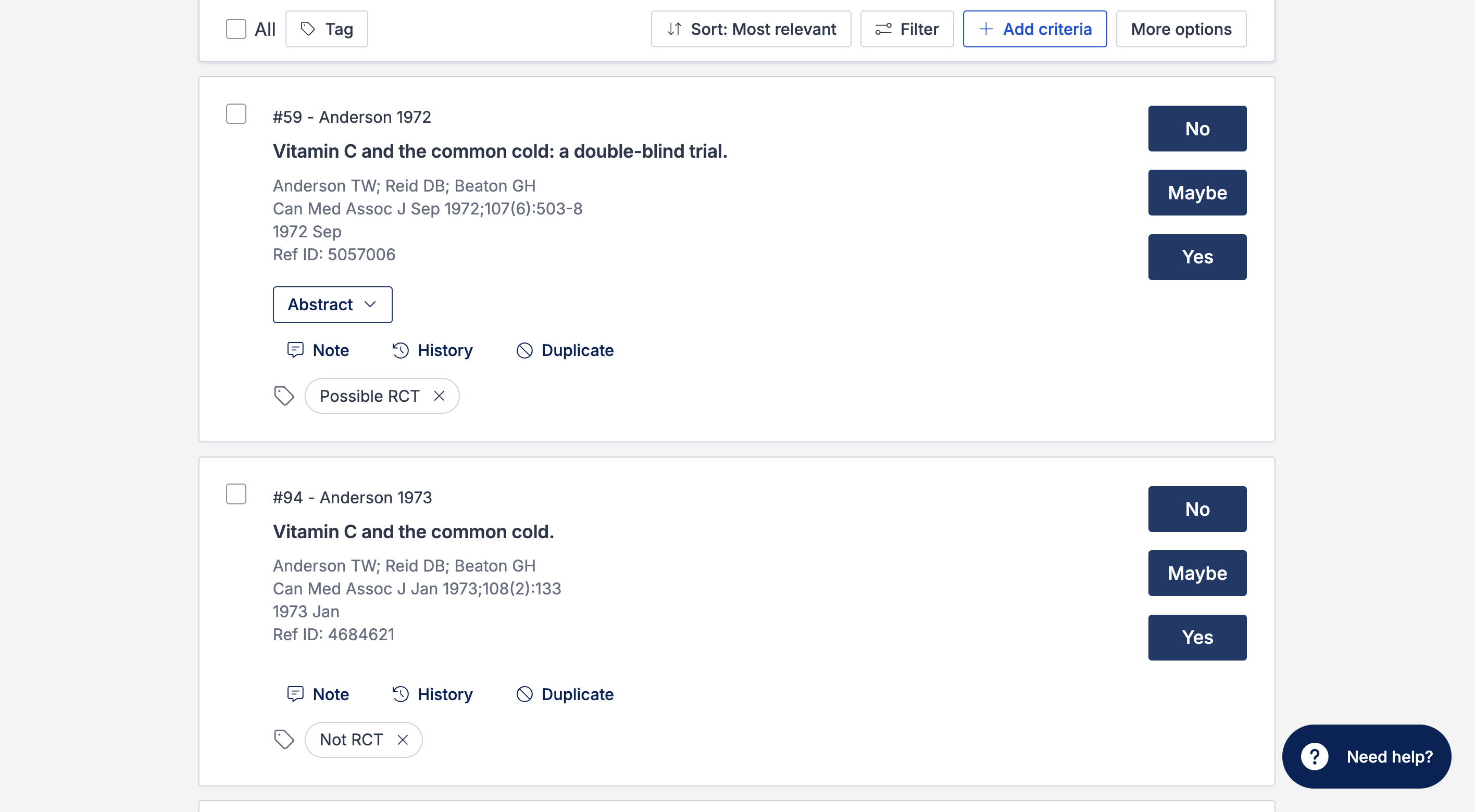
The studies list can also be filtered by the RCT tags, allowing you to see a list of only “Possible RCTs” or “Not RCTs”:
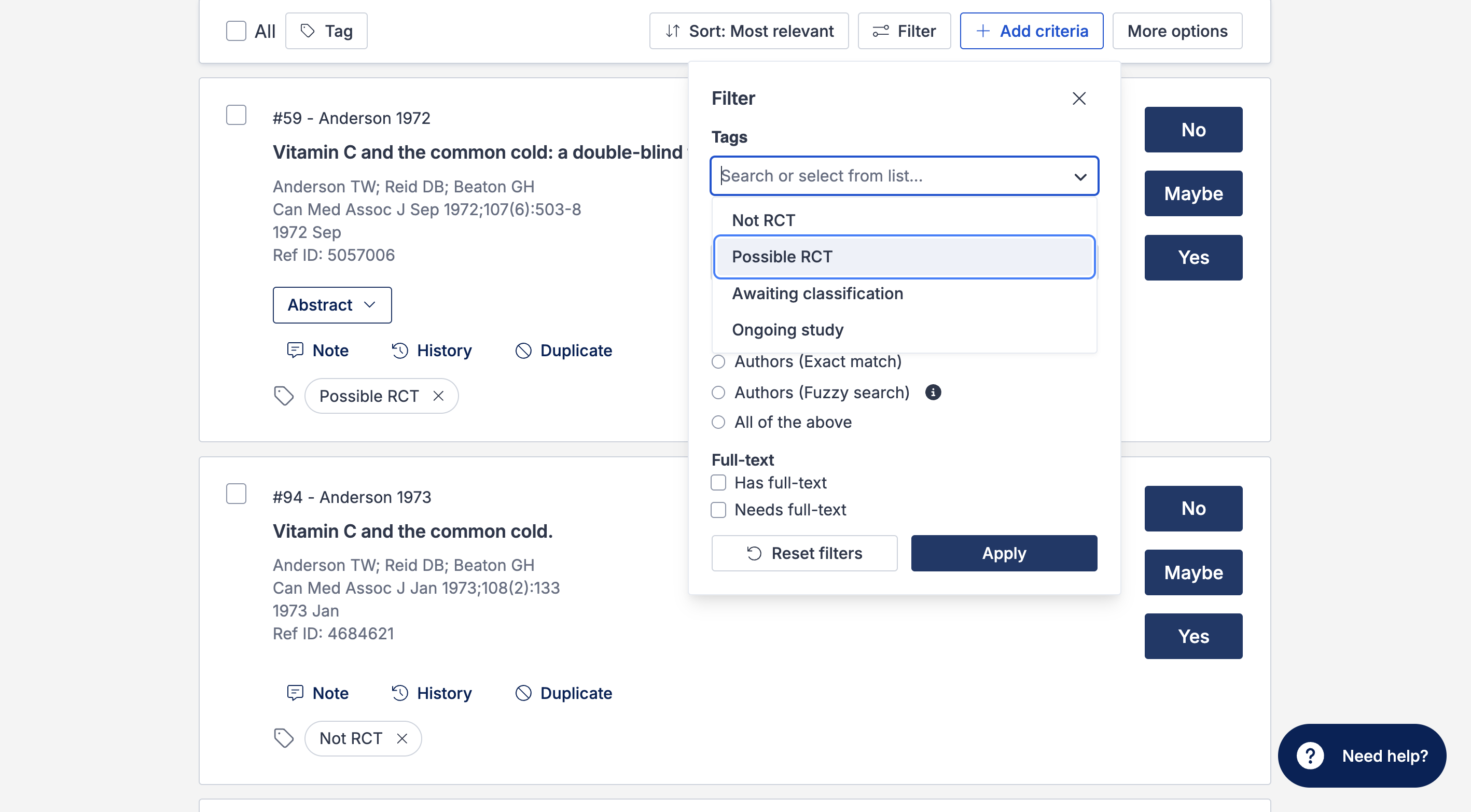
The Cochrane RCT Classifier
We’ve integrated with the leading RCT classifier algorithm, developed by EPPI-Centre, to provide you with a prediction on whether studies in your review potentially report on an RCT. The Classifier has been endorsed by Cochrane and is also being used in the Cochrane Screen4Me workflows, which also includes Cochrane Crowd.
Evaluation
The classifier has been shown in testing to successfully identify over 99.5% of health-related references that potentially report on RCTs, only incorrectly classifying ~0.5% of RCTs as not being an RCT (Thomas et al, 2021).
Known limitations
Given the RCT classifier has been trained, calibrated and validated using health-related research papers, we only allow the features to be used on Cochrane reviews or reviews in the “medical and health science” research area.
To ensure the classifier has enough context about each reference to make informed predictions, we'll only input references with titles of 14 characters or more and abstracts of 400 characters or more. This mirrors the criteria used for validating the classifier's performance.
Enabling the feature
When creating a review, you will be shown RCT-related settings under the “Automation options” section when the review’s research area is set to “Medical and health sciences” or it is a Cochrane review:
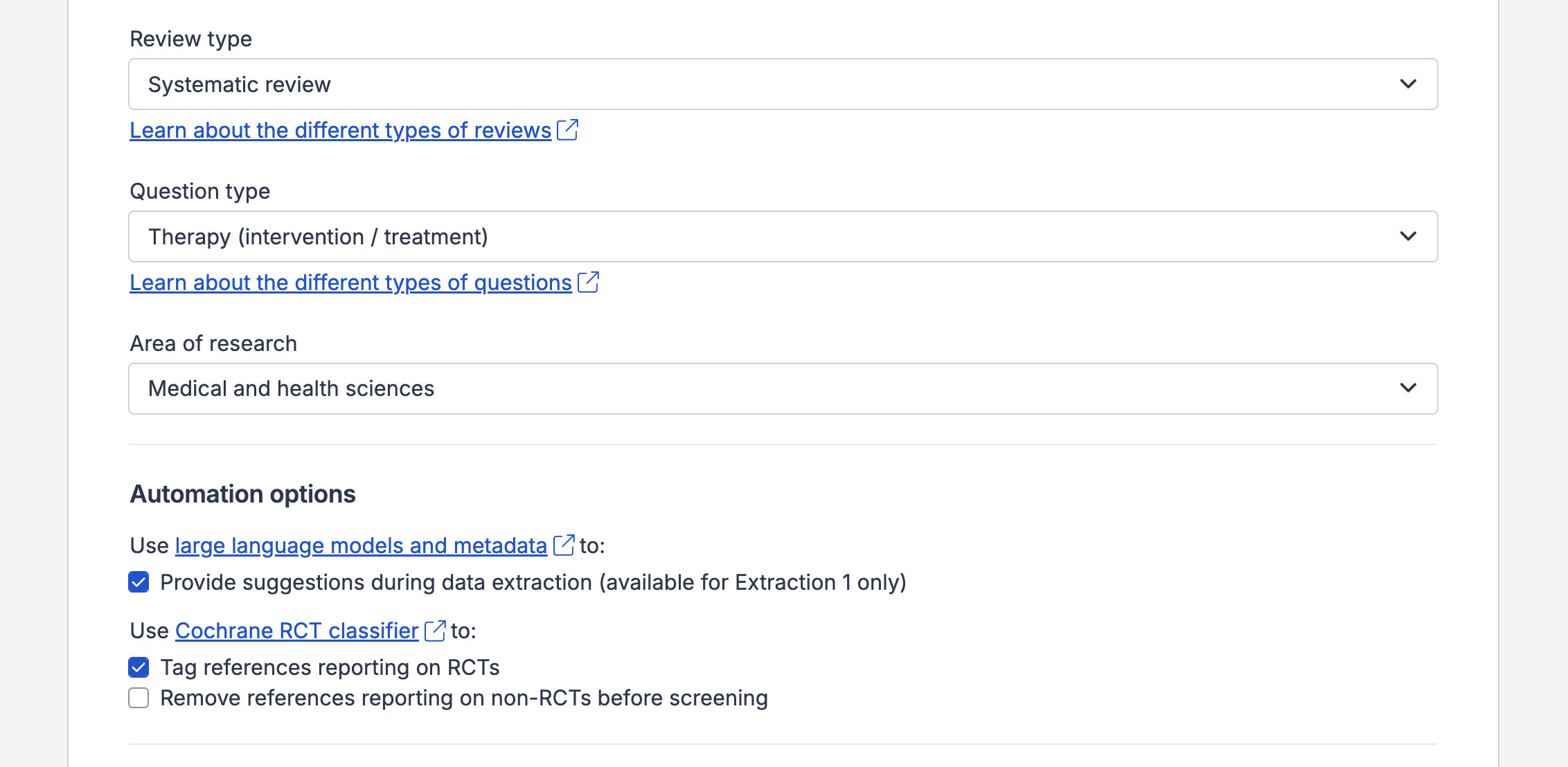
Disabling the feature
You can disable the feature at any point via the automation options section on review settings (shown above).
Disabling the feature will retain tags already applied to references, although further references won’t be automatically tagged when imported to the review.
Reporting feature usage
For your Manuscript (Methods Section) use the following text to transparently report use of this feature in line with RAISE standards:
We will use the “Tagging references reporting on RCTs” feature (version number not supplied, accessed on dd/mm/yyyy) developed by Covidence for tagging references that are likely to describe possible randomized controlled trials and those that are not likely to describe randomized controlled trials in the Title & Abstract screening stage. The tool will be used with no customisation, training or parameter changes applied.
Outputs from the tool are justified for use in our synthesis because:
Human reviewers retain full control: All references, tagged or untagged, will be screened by human reviewers at both the Title & Abstract and Full Text stages, ensuring that human judgment determines all study selection decisions.
High-sensitivity model performance: The classifier, developed by the EPPI-Centre and endorsed by Cochrane, identifies more than 99.5% of health-related references that potentially report RCTs and misclassifies only ~0.5% of true RCTs as non-RCTs (Thomas et al., 2021), prioritising high sensitivity to minimise false exclusions.
Mitigation for short or uninformative abstracts: A known limitation of the classifier is reduced performance on records with very short or poorly reported titles/abstracts. Covidence mitigates this risk by only attempting to apply the RCT/non-RCT tag to records that meet minimum information thresholds (title >14 characters and abstract >400 characters), consistent with the filters used in the original evaluation. Records below these thresholds are deliberately left untagged and are screened entirely by human reviewers, ensuring that low-information records receive full manual assessment.
Limitations of the tool include:
Evaluation limitations: The evaluation by Thomas et al. (2021) had limitations that users of this feature should be aware of. The classifier was trained and evaluated primarily on English-language, biomedical records (Embase, Cochrane, Clinical Hedges), so performance in other languages, non-health domains, or niche subfields is less certain. Its best performance (99.5% recall) is observed only for records with sufficiently long and informative titles/abstracts; older studies and very short or poorly reported abstracts are more likely to be misclassified and require manual handling. Because the classifier analyses only the title and abstract, and does not use full text or additional metadata, any signals present only in the full text will not be detected.
Risk of automation bias: While all references are still assessed by human reviewers, the presence of misclassified tags may influence their independent judgment in ways the tool cannot fully safeguard against.

